Sensitivity of Coastal Environments and Wildlife to Spilled Oil: Florida Panhandle: HABITATS (Habitat Polygons)
Data Set (DS) | Office of Response and Restoration (ORR)GUID: gov.noaa.nmfs.inport:40716 | Updated: May 15, 2025 | Published / External
- View As
- View in Hierarchy
Summary
Short Citation
Office of Response and Restoration, 2025: Sensitivity of Coastal Environments and Wildlife to Spilled Oil: Florida Panhandle: HABITATS (Habitat Polygons), https://www.fisheries.noaa.gov/inport/item/40716.
Full Citation Examples
This data set contains sensitive biological resource data for rare plants for the Florida Panhandle. Vector polygons in this data set represent rare plant occurrences. Species specific abundance, seasonality, status, life history, and source information are stored in relational data tables (described below) designed to be used in conjunction with this spatial data layer. This data set comprises a portion of the ESI data for the Florida Panhandle. ESI data characterize the marine and coastal environments and wildlife by their sensitivity to spilled oil. The ESI data include information for three main components: shoreline habitats, sensitive biological resources, and human-use resources.
PurposeThe ESI data were collected, mapped, and digitized to provide environmental data for oil spill planning and response. The Clean Water Act with amendments by the Oil Pollution Act of 1990 requires response plans for immediate and effective protection of sensitive resources.
Distribution Information
-
Downloadable Data
None
DO NOT USE MAPS FOR NAVIGATIONAL PURPOSES.
Besides the above warning, there are no use constraints on these data. Note that the ESI database should not be used to the exclusion of other pertinent data or information held by state or federal agencies or other organizations. Likewise, information contained in the database cannot be used in place of consultations with environmental, natural resource, and cultural resource agencies, or in place of field surveys. Recognize that the information contained in the ESI database represents known concentration areas or occurrences of natural, cultural, and human-use resources, but does not necessarily represent the full distribution or range of each species or resource. This is particularly important to recognize when considering potential impacts to protected resources, such as endangered species, wetlands, etc. Acknowledgment of the originators, publishers, contributors, and sources listed would be appreciated in products derived from these data.
Controlled Theme Keywords
biota, environment
URLs
Contact Information
Point of Contact
ESI Program Manager
orr.esi@noaa.gov
Metadata Contact
ESI Program Manager
orr.esi@noaa.gov
Extents
-87.625° W,
-83.684° E,
30.747° N,
28.277° S
2007 - 2011
Item Identification
| Title: | Sensitivity of Coastal Environments and Wildlife to Spilled Oil: Florida Panhandle: HABITATS (Habitat Polygons) |
|---|---|
| Short Name: | flpan_habitats |
| Status: | Completed |
| Publication Date: | 2012-08 |
| Abstract: |
This data set contains sensitive biological resource data for rare plants for the Florida Panhandle. Vector polygons in this data set represent rare plant occurrences. Species specific abundance, seasonality, status, life history, and source information are stored in relational data tables (described below) designed to be used in conjunction with this spatial data layer. This data set comprises a portion of the ESI data for the Florida Panhandle. ESI data characterize the marine and coastal environments and wildlife by their sensitivity to spilled oil. The ESI data include information for three main components: shoreline habitats, sensitive biological resources, and human-use resources. |
| Purpose: |
The ESI data were collected, mapped, and digitized to provide environmental data for oil spill planning and response. The Clean Water Act with amendments by the Oil Pollution Act of 1990 requires response plans for immediate and effective protection of sensitive resources. |
| Notes: |
1726 |
| Other Citation Details: |
Prepared by Research Planning, Inc., Columbia, South Carolina for the National Oceanic and Atmospheric Administration (NOAA), National Ocean Service, Office of Response and Restoration, Emergency Response Division, Seattle, Washington. |
Keywords
Theme Keywords
| Thesaurus | Keyword |
|---|---|
| ISO 19115 Topic Category |
biota
|
| ISO 19115 Topic Category |
environment
|
| UNCONTROLLED | |
| NOS Data Explorer Topic Category | Environmental Monitoring |
| None | Coastal resources |
| None | Coastal Zone Management |
| None | Environmental Monitoring |
| None | ESI |
| None | Habitat |
| None | Oil spill planning |
| None | Sensitivity maps |
| None | Wildlife |
Spatial Keywords
| Thesaurus | Keyword |
|---|---|
| UNCONTROLLED | |
| None | Florida Panhandle |
Physical Location
| Organization: | Office of Response and Restoration |
|---|---|
| City: | Silver Spring |
| State/Province: | MD |
Data Set Information
| Data Set Scope Code: | Data Set |
|---|---|
| Maintenance Frequency: | None Planned |
| Data Presentation Form: | vector digital data |
| Entity Attribute Overview: |
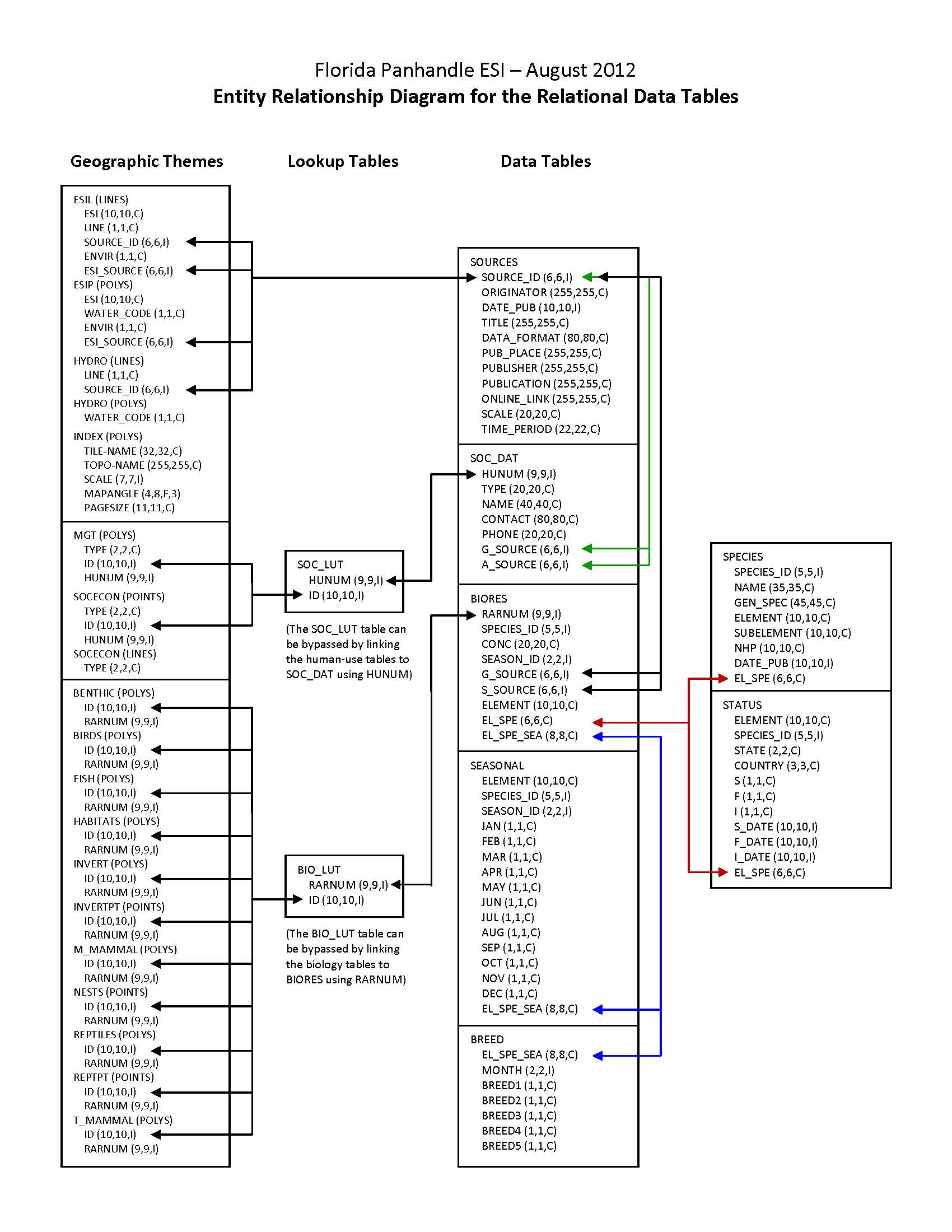
In addition to the geographic data layers, six relational attribute or data tables (BIORES, BREED, SEASONAL, SOURCES, SPECIES, and STATUS) are used to store the complex biological data in the ESI data structure. The geographic data layer containing biological resource information (in this case, HABITATS) is linked to the Biological Resources table (BIORES) using the unique ID and the lookup table BIO_LUT, or it can be linked directly using RARNUM. The ID is a unique combination of the atlas number (for the Florida Panhandle atlas, the number is 218), an element/layer specific number (BIRDS are layer 1, FISH are layer 2, etc.), and a unique record number. The RARNUM represents a unique combination of species, seasonalities, concentrations, and source information. For each of these groupings, a number is generated. That number is concatenated with the atlas number to create a "resource at risk" number that is unique across atlases. BIORES and the other relational data tables are described in the Detailed_Description sections. See the Browse_Graphic section for a link to the entity-relationship diagram, which describes the way these tables relate to the geographic data layers and other attribute tables in the ESI data structure. Due to the complexity of the relational database model, the data items are also post-processed into a flat file format. This table, called BIOFILE, may be used in place of the relational files to ease simple data queries. The items in the flat file are ELEMENT, SUBELEMENT, NAME, GEN_SPEC, S, F, NHP, DATE_PUB, CONC, JAN, FEB, MAR, APR, MAY, JUN, JUL, AUG, SEP, OCT, NOV, DEC, BREED1, BREED2, BREED3, BREED4, BREED5, RARNUM, G_SOURCE, S_SOURCE, and BREED. All of these items are the same as their counterparts in the individual data tables (described in the Detailed_Description sections), except the BREED1-BREED5 and BREED items. BREED is a newly generated variable used to link to the BREED_DT data table, a modified, more compact version of the relational BREED data table. BREED1-BREED5 give a text summary of when each life stage occurs within the associated map object. The life stages referred to are the same as those listed in the Detailed_Description of the BREED data table. The link to the BIOFILE may be made through the BIO_LUT, using ID to link to RARNUM, or BIOFILE may be linked directly to the RARNUM in each of the geographic layer's attribute data tables. As mentioned, BREED_DT is an auxiliary support data table to the flat file structure, which allows the user to do searches based on month for seasonal breeding activities. The link from the flat file to BREED_DT is the BREED item. A second supporting data table is SOURCES. This is the same as the source file described above, and the link from the flat file is both G_SOURCE and S_SOURCE. It should be noted that although the flat file eases data query, it is not a normalized database structure, and actual updates performed by the states and other responsible agencies should be done using the relational data tables. The entity-relationship diagram, describing relationships between attribute tables in the ESI data structure does NOT include the BIOFILE data table, and this data table is NOT described in a Detailed_Description section. |
| Entity Attribute Detail Citation: |
A complete description of entity types, attributes, and attribute values for ESI atlases can be found in the NOAA ESI Guidelines (http://response.restoration.noaa.gov/esi_guidelines). |
| Distribution Liability: |
These data represent a snapshot in time and temporal changes may have occurred. These data are not intended to include all biological or human-use resources present in an area; they focus on species and resources particularly sensitive to oiling. In the event of a spill, they should be used for a first assessment only. The data providers are the experts with regard to individual resources. They should be contacted to confirm if more current data exist, and/or in-depth information is needed about a particular resource. |
| Data Set Credit: | This project was supported by the National Oceanic and Atmospheric Administration (NOAA), National Ocean Service (NOS), Office of Response and Restoration (OR&R), Emergency Response Division (ERD), Seattle, Washington; the Department of Homeland Security (DHS), United States Coast Guard (USCG), Office of Incident Management and Preparedness Washington, D.C.; and the Fish and Wildlife Research Institute (FWRI), Florida Fish and Wildlife Conservation Commission, St. Petersburg, Florida. |
Support Roles
Data Steward
| Date Effective From: | 2012-08 |
|---|---|
| Date Effective To: | |
| Contact (Position): | ESI Program Manager |
| Address: |
7600 Sand Point Way NE Seattle, WA 98115 |
| Email Address: | orr.esi@noaa.gov |
Distributor
| Date Effective From: | 2012-08 |
|---|---|
| Date Effective To: | |
| Contact (Position): | ESI Program Manager |
| Address: |
7600 Sand Point Way NE Seattle, WA 98115 |
| Email Address: | orr.esi@noaa.gov |
Metadata Contact
| Date Effective From: | 2012-08 |
|---|---|
| Date Effective To: | |
| Contact (Position): | ESI Program Manager |
| Address: |
7600 Sand Point Way NE Seattle, WA 98115 |
| Email Address: | orr.esi@noaa.gov |
Point of Contact
| Date Effective From: | 2012-08 |
|---|---|
| Date Effective To: | |
| Contact (Position): | ESI Program Manager |
| Address: |
7600 Sand Point Way NE Seattle, WA 98115 |
| Email Address: | orr.esi@noaa.gov |
Extents
| Currentness Reference: | The data were compiled during 2010-2012. The currentness dates for the data range from 2007 to 2011 and are documented in the Lineage section. |
|---|
Extent Group 1
Extent Group 1 / Geographic Area 1
| W° Bound: | -87.625 | |
|---|---|---|
| E° Bound: | -83.684 | |
| N° Bound: | 30.747 | |
| S° Bound: | 28.277 |
Extent Group 1 / Time Frame 1
| Time Frame Type: | Range |
|---|---|
| Start: | 2007 |
| End: | 2011 |
Spatial Information
Spatial Representation
Representations Used
| Vector: | Yes |
|---|
Vector Representation 1
| Complex Object Present?: | Yes |
|---|---|
| Complex Object Count: | 3018 |
| Curve Object Present?: | Yes |
| Curve Object Count: | 97974 |
| Point Object Present?: | Yes |
| Point Object Count: | 3019 |
| Surface Object Present?: | Yes |
| Surface Object Count: | 2968 |
Access Information
| Security Class: | Unclassified |
|---|---|
| Data Access Procedure: |
Contact NOAA for distribution options (see Distributor). ESI data are processed into multiple formats. Distribution formats include a Geodatabase (including an ArcMap .mxd file, complete with database links and symbology), ARC export files, and shapefiles. The database files, available in text and INFO(R) formats, are provided in both the NOAA standard relational database format (see NOAA Technical Memorandum NOS ORCA 115) and in a simplified desktop flat file format. This metadata document includes information about both of these database formats.; |
| Data Access Constraints: |
None |
| Data Use Constraints: |
DO NOT USE MAPS FOR NAVIGATIONAL PURPOSES. Besides the above warning, there are no use constraints on these data. Note that the ESI database should not be used to the exclusion of other pertinent data or information held by state or federal agencies or other organizations. Likewise, information contained in the database cannot be used in place of consultations with environmental, natural resource, and cultural resource agencies, or in place of field surveys. Recognize that the information contained in the ESI database represents known concentration areas or occurrences of natural, cultural, and human-use resources, but does not necessarily represent the full distribution or range of each species or resource. This is particularly important to recognize when considering potential impacts to protected resources, such as endangered species, wetlands, etc. Acknowledgment of the originators, publishers, contributors, and sources listed would be appreciated in products derived from these data. |
Distribution Information
Distribution 1
| Download URL: | https://response.restoration.noaa.gov/esi_download |
|---|---|
| Distributor: | |
| Description: |
Downloadable Data |
| File Type (Deprecated): | Multiple formats |
URLs
URL 1
| URL: | https://response.restoration.noaa.gov/esi |
|---|---|
| URL Type: |
Online Resource
|
URL 2
| URL: | https://response.restoration.noaa.gov/esi_download |
|---|---|
| URL Type: |
Online Resource
|
URL 3
| URL: | https://response.restoration.noaa.gov/esi_guidelines |
|---|---|
| URL Type: |
Online Resource
|
URL 4
| URL: | https://response.restoration.noaa.gov/sites/default/files/esimaps/gisdata/FloridaPanhdle_2012_datafig.jpg |
|---|---|
| URL Type: |
Browse Graphic

|
| File Resource Format: | JPEG |
| Description: |
Depicts the relationships between spatial data layers and attribute data tables for the Florida Panhandle ESI data. |
URL 5
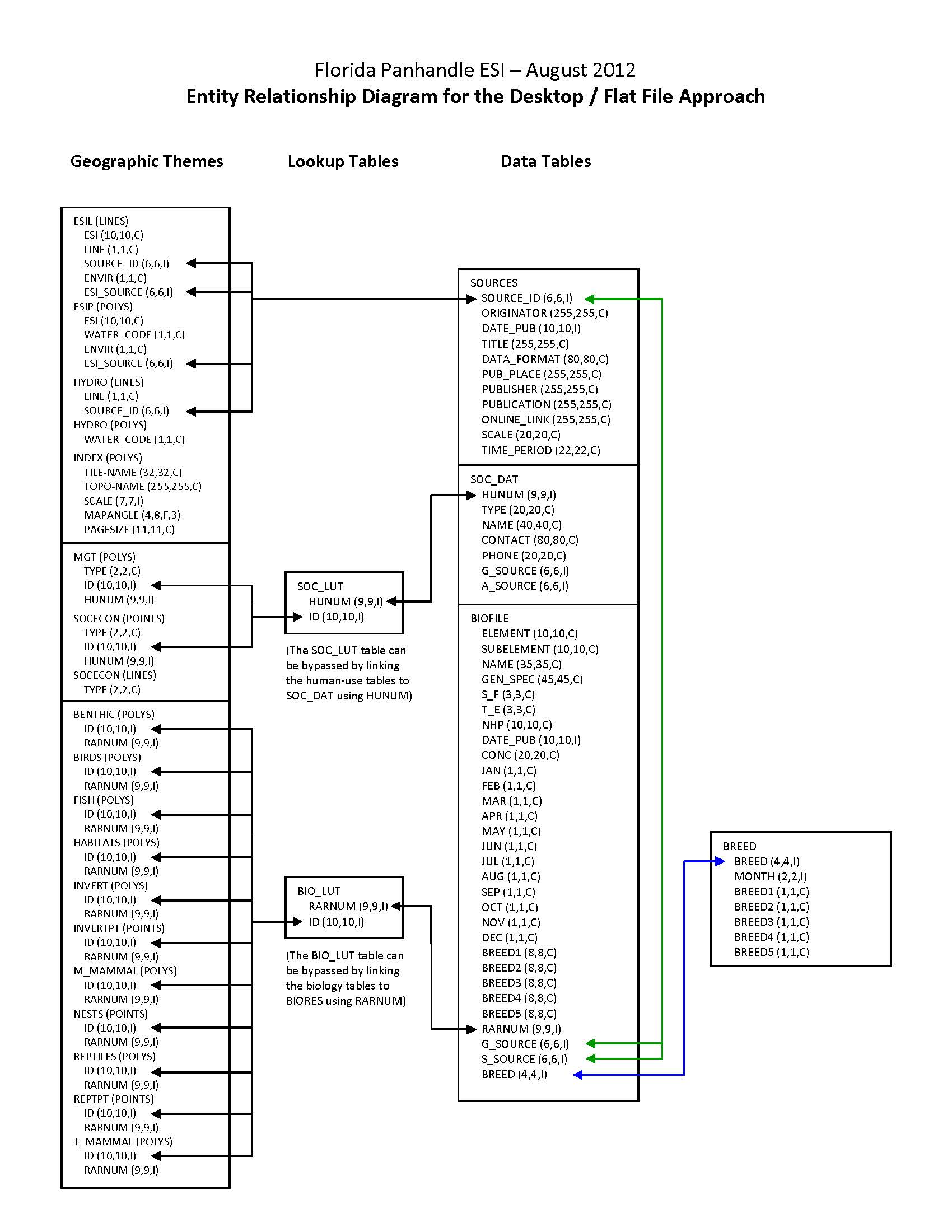
| URL: | https://response.restoration.noaa.gov/sites/default/files/esimaps/gisdata/FloridaPanhdle_2012_datafig2.jpg |
|---|---|
| URL Type: |
Browse Graphic
|
| File Resource Format: | JPEG |
| Description: |
Depicts the relationships between spatial data layers and desktop data tables for the Florida Panhandle ESI data. |
Activity Log
Activity Log 1
| Activity Date/Time: | 2014-06-09 |
|---|---|
| Description: |
Date that the source FGDC record was last modified. |
Activity Log 2
| Activity Date/Time: | 2017-04-05 |
|---|---|
| Description: |
Converted from Content Standards for Digital Geospatial Metadata (version FGDC-STD-001-1998) using 'fgdc_to_inport_xml.pl' script. Contact Tyler Christensen (NOS) for details. |
Activity Log 3
| Activity Date/Time: | 2017-09-13 |
|---|---|
| Description: |
Partial upload of Spatial Info section only. |
Activity Log 4
| Activity Date/Time: | 2017-11-01 |
|---|---|
| Description: |
Replaced entire Lineage section to populate new Source Contribution field. |
Activity Log 5
| Activity Date/Time: | 2018-02-08 |
|---|---|
| Description: |
Partial upload of Positional Accuracy fields only. |
Technical Environment
| Description: |
The software packages used to develop the atlas are Environmental Systems Research Institute's ARC/INFO(R) (version 9.3) and SQL SERVER(R) (version 2000). The hardware configuration is PCs with Windows Operating System (2000/XP/2003). The Spatial_Data_Organization Information section refers only to the source files in the ARC export format. The following files are included in the data set: benthic.e00, birds.e00, esil.e00, esip.e00, fish.e00, habitats.e00, hydro.e00, index.e00, invert.e00, invertpt.e00, m_mammal.e00, mgt.e00, nests.e00, reptiles.e00, reptpt.e00, socecon.e00, and t_mammal.e00. Associated relational and desktop data tables provided in Arc export and text format are bio_lut.e00, biofile.e00, biores.e00, breed.e00, breed_dt.e00, seasonal.e00, soc_dat.e00, soc_lut.e00, sources.e00, species.e00, and status.e00. |
|---|
Data Quality
| Accuracy: |
A multi-stage error checking process is used to verify both attribute accuracy and logical consistency throughout data production. The process includes a standardized data entry methodology, hardcopy data review by in-house and external resource experts, a final Quality Assurance/Quality Control (QA/QC) process, and multiple automated logical consistency checks. Quantitative data (such as densities, counts, abundances, or concentrations) provided by resource experts for inclusion in the data set may vary widely in attribute accuracy, depending upon the methodology used to collect and compile such data. For a more detailed evaluation of source data attribute accuracy, contact the sources listed in the Lineage section. |
|---|---|
| Horizontal Positional Accuracy: |
Spatial components for the biological data layers can come from expert interviews, hardcopy, or digital sources. Some of the spatial components of the biological data layers may have been developed using regional experts who estimate concentration areas. It is difficult to estimate the positional accuracy of such data, except to state that they are compiled on hardcopy base maps with a scale of 1:24,000. Some of the spatial components of the biological data sets are developed from pre-existing digital or hardcopy sources and reflect the positional accuracy of these original data. Note that biological resource data by their very nature are considered "fuzzy", and this should be understood when considering the positional accuracy of vector digital objects representing these resources. See the Lineage and Process_Description sections for more information on the original source data and how these data were integrated or manipulated to create the final data set. |
| Completeness Report: |
These data represent a synthesis of digital data sets on rare plant occurrences. These data do not necessarily represent all habitat occurrences in Florida Panhandle. The following species are included in this data set: (Species_ID, Common Name, Scientific Name [n/a if not applicable]): 228, Chapman's sedge, Carex chapmanii; 241, Bumpy jointtail grass, Mnesithea tuberculosa; 242, Pond spice, Litsea aestivalis; 246, Southern milkweed, Asclepias viridula; 663, Beaked spikerush, Eleocharis rostellata; 684, Violet butterwort, Pinguicula ionantha; 796, Manyflower grasspink, Calopogon multiflorus; 800, Godfrey's goldenaster, Chrysopsis godfreyi; 813, Sarvis holly, Ilex amelanchier; 815, Panhandle lily, Lilium iridollae; 816, Bog spicebush, Lindera subcoriacea; 820, Flameflower, Macranthera flammea; 830, Largeleaf jointweed, Polygonella macrophylla; 832, Florida pondweed, Potamogeton floridanus; 836, Mosquito beadsedge, Rhynchospora crinipes; 840, Nightflowering wild petunia, Ruellia noctiflora; 842, Georgia bully, Sideroxylon thornei; 845, Harper's yelloweyed grass, Xyris scabrifolia; 848, Crimson pitcherplant, Sarracenia leucophylla; 952, Perforate reindeer lichen, Cladonia perforata; 953, Pinewoods bluestem, Andropogon arctatus; 955, Florida wild indigo, Baptisia calycosa var. villosa; 956, Apalachicola aster, Eurybia spinulosa; 957, Scareweed, Baptisia simplicifolia; 958, Flyr's nemesis, Brickellia cordifolia; 959, Florida calamint, Clinopodium dentatum; 960, Florida sandreed, Calamovilfa curtissii; 961, Eastern sweetshrub, Calycanthus floridus; 962, Baltzell's sedge, Carex baltzellii; 963, Cruise's goldenaster, Chrysopsis gossypina ssp. Cruiseana; 965, Tropical waxweed, Cuphea aspera; 966, Threadleaf sundew, Drosera filiformis; 967, Spoonleaf sundew, Drosera intermedia; 968, Florida burhead, Echinodorus floridanus; 969, Trailing arbutus, Epigaea repens; 970, Blackbract pipewort, Eriocaulon nigrobracteatum; 971, Telephus spurge, Euphorbia telephioides; 972, Wiregrass gentian, Gentiana pennelliana; 974, Godfrey's spiderlily, Hymenocallis godfreyi; 975, Henry's spiderlily, Hymenocallis henryae; 976, Smoothbark St. Johnswort, Hypericum lissophloeus; 977, Pennsylvania rush, Juncus gymnocarpus; 978, Thickleaf water-willow, Justicia crassifolia; 979, Mountain laurel, Kalmia latifolia; 980, Corkwood, Leitneria floridana; 981, Kral's yelloweyed grass, Xyris longisepala; 982, Petiteplant, Lepuropetalon spathulatum; 983, Godfrey's blazing star, Liatris provincialis; 984, West's flax, Linum westii; 985, Gulf Coast lupine, Lupinus westianus; 986, Curtiss' loosestrife, Lythrum curtissii; 987, White birds-in-a-nest, Macbridea alba; 988, Ashe's magnolia, Magnolia ashei; 989, Pyramid magnolia, Magnolia pyramidata; 990, Green adder's-mouth orchid, Malaxis unifolia; 991, Alabama milkvine, Matelea alabamensis; 992, Pinesap, Monotropa hypopithys; 993, Florida beargrass, Nolina atopocarpa; 994, Giant cowbane, Oxypolis greenmanii; 995, Naked-stemmed panicgrass, Dichanthelium nudicaule; 996, Carolina grass of Parnassus, Parnassia caroliniana; 997, Paper nailwort, Paronychia chartacea var. minima; 998, Pineland false sunflower, Phoebanthus tenuifolius; 1000, Godfrey's false dragonhead, Physostegia godfreyi; 1059, Southern butterwort, Pinguicula primuliflora; 1060, Zigzag silkgrass, Pityopsis flexuosa; 1061, Small green wood orchid, Platanthera clavellata; 1062, Arkansas oak, Quercus arkansana; 1063, White meadowbeauty, Rhexia parviflora; 1064, Panhandle meadowbeauty, Rhexia salicifolia; 1065, Orange azalea, Rhododendron austrinum; 1066, Sweet pitcherplant, Sarracenia rubra; 1067, Bay starvine, Schisandra glabra; 1068, Florida skullcap, Scutellaria floridana; 1069, Mock pennyroyal, Stachydeoma graveolens; 1070, Silky camellia, Stewartia malacodendron; 1071, Pineland hoarypea, Tephrosia mohrii; 1072, Cooley's meadow-rue, Thalictrum cooleyi; 1073, Chapman's crownbeard, Verbesina chapmanii; 1074, Quillwort yelloweyed grass, Xyris isoetifolia. |
| Conceptual Consistency: |
A multi-stage error checking process, described in the above Attribute_Accuracy_Report, is used to verify both attribute accuracy and logical consistency throughout data production. This process includes multiple automated logical consistency checks that test the files for missing or duplicate data, rules for proper coding, GIS topological consistencies (such as dangles, unnecessary nodes, etc.), and SQL SERVER(R) to ARC/INFO(R) consistencies. After the data are delivered to NOAA, they are again subjected to a number of quality and consistency checks. In the process of checking for topological and database consistencies, new IDs and RARNUMs or HUNUMs are also generated. The new IDs are a combination of atlas number, element number, and record number. In addition, the value used to represent the element is modified to reflect the type of feature being mapped. In the case of an element that is normally represented by a point or polygon, a value of 20 is added to the standard element value for mapping of linear features. In the case where an element usually mapped as a polygon is represented by a point, a value of 30 is added to the regular element value. The RARNUMs are also modified to include the atlas number, so multiple atlases can be combined and RARNUMs remain unique. RARNUMs are redefined on an element basis, so "resource at risk" groupings will contain only a single element. HUNUMs are also modified to include the atlas number. |
Lineage
Sources
EGLIN CLADONIA PERFORATA HAB AREA
| Contact Name: | USAF EGLIN AIR FORCE BASE |
|---|---|
| Publish Date: | 2007-01-01 |
| Extent Type: | Discrete |
| Extent Start Date/Time: | 2007 |
| Source Contribution: |
HABITATS INFORMATION | Source Geospatial Form: vector digital data | Type of Source Media: EMAIL |
ELEMENT OCCURRENCE POLYGON DATA LAYER
| Contact Name: | FLORIDA NATURAL AREAS INVENTORY (FNAI) |
|---|---|
| Publish Date: | 2011-01-01 |
| Extent Type: | Discrete |
| Extent Start Date/Time: | 2011 |
| Source Contribution: |
HABITATS INFORMATION | Source Geospatial Form: vector digital data | Type of Source Media: EMAIL |
Process Steps
Process Step 1
| Description: |
The main sources of data used to depict habitat distribution and seasonality for this data layer were digital data sets provided by Florida Fish and Wildlife Conservation Commission-Fish and Wildlife Research Institute (FWC-FWRI), Eglin Air Force Base, and Florida Natural Areas Inventory (FNAI). The above digital and/or hardcopy sources were compiled by the project biologist to create the HABITATS data layer. Depending on the type of source data, three general approaches are used for compiling the data layer: 1) information gathered during initial interviews and from hardcopy sources are compiled onto U.S. Geological Survey 1:24,000 topographic quadrangles and digitized; 2) hardcopy maps are digitized at their source scale; 3) digital data layers are evaluated and used "as is" or integrated with the hardcopy data sources. See the Lineage section for additional information on the type of source data for this data layer. The compiled ESI, biology, and human-use data are plotted onto hardcopy draft maps. Following the delivery of draft maps to the participating resource experts, a second set of interviews are conducted to review the maps. If necessary, edits to the HABITATS data layer are made based on the recommendations of the resource experts, and final hardcopy maps and digital data are created. |
|---|---|
| Process Date/Time: | 2012-08-01 00:00:00 |
Child Items
Catalog Details
| Catalog Item ID: | 40716 |
|---|---|
| GUID: | gov.noaa.nmfs.inport:40716 |
| Metadata Record Created By: | Tyler Christensen |
| Metadata Record Created: | 2017-04-05 14:53+0000 |
| Metadata Record Last Modified By: | SysAdmin InPortAdmin |
| Metadata Record Last Modified: | 2025-05-15 19:16+0000 |
| Metadata Record Published: | 2018-02-08 |
| Owner Org: | ORR |
| Metadata Publication Status: | Published Externally |
| Do Not Publish?: | N |
| Metadata Last Review Date: | 2018-02-08 |
| Metadata Review Frequency: | 1 Year |
| Metadata Next Review Date: | 2019-02-08 |
