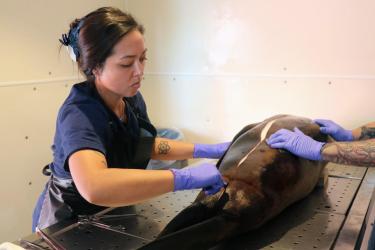
Cali Turner Tomaszewicz has worked with the Southwest Fisheries Science Center Marine Turtle Ecology and Assessment Program since 2008. Her current research uses humerus bones of dead, stranded sea turtles and applies stable isotope analysis with skeletochronology to focus on their life history, habitat use, and growth patterns. Her most recent published work focuses on East Pacific green sea turtles and North Pacific loggerhead turtles.
How did you first become involved in sea turtle research?
I grew up in Colorado, so nature and the outdoors were always my thing. But my grandparents lived in Florida and there was a sea turtle rehabilitation facility near their house. We always went there when I was a kid so sea turtles had a special place in my heart—they are fascinating animals! I had also worked as a marine conservation educator and I really loved teaching about the ocean and showing people how ocean science is relevant to their everyday lives.
After teaching for a few years, I was ready to conduct some of my own scientific research. So in 2008, I started my master’s program at Scripps Institution of Oceanography and through that program I started working with Jeff Seminoff. We were studying turtles near a power plant in San Diego Bay that was being shut down, and I just got hooked on doing research and working with the Marine Turtle Ecology team. I loved the applied aspect of sea turtle research, and because turtles are such charismatic animals they are a great way to begin talking to people about other important issues related to ocean health and sustainable living. Between my master’s degree and Ph.D., I worked on my first paper with Jeff. Because I wanted to continue doing research, we came up with the plan to explore and develop a new approach combining skeletochronology and precise sequential isotope analysis.
You helped develop this new technique to examine sea turtle life history in finer detail. Can you tell us more about that process?

Skeletochronology has been used for more than 30 years as a way to estimate the age of sea turtles—it’s just like using the rings of a tree. Larisa Avens had been doing skeletochronology research and analyzing age and growth patterns in sea turtles, and she and Lisa Goshe at the NOAA Beaufort Lab in North Carolina trained me on how to process bones for skeletochronology. Also, earlier work by Melissa Snover had started exploring the patterns of stable isotope values in turtle bones to detect a shift from offshore to nearshore habitats. My task was to refine the process of combining these two techniques in order to recreate a detailed past life history for turtles. This had never been done at this level for any turtles, and so our next step was to try to make the process really precise to extract information about diet, growth, and more from the individual bone growth layers.
Together with Larisa, Jeff, and many more collaborators, I wrote up a full protocol on how to best combine these two techniques; we identified tips, tricks, and challenges, and this work resulted in three methods-based papers for my Ph.D. We then applied these methods to two populations of turtles in the Pacific (green turtles and loggerheads), which led to three more papers for my Ph.D. Those papers revealed some interesting facts about where these turtles spent their time, how long it took them to reach maturity, and how not all the turtles in a single population do the same things—there is a lot of variability in behavior. The findings from those papers have been useful for management, because a key focus of the work was figuring out how much these turtles are using particular habitats off the coast of a site in Baja, Mexico, where there can be a lot of overlap with fishing.
What were some of the biggest challenges and/or surprises in developing this new approach?
I was surprised that there wasn’t already a protocol for this. Through the process I’ve learned to appreciate that is the nature of doing something new and breaking new ground. Getting over that expectation that there must already be a way to do this, and then digging in and figuring it out on your own (or, in my case, with a good team) was the really fun part of this project. We had to determine the smallest details, like the angle of the drill, how to align and “map” where the different growth layers were, and how to extract samples from very small growth layers. We played around with different ways of sampling sections. For instance, the drill takes hours to take a sample, so we tested to see if a biopsy core sample could also work. It was faster but it didn’t work as well; the isotope values were not reliable.
Putting all the data together to get these multiyear ecological trends was another challenge. We can look at 20 years of growth, size, age, and diet for one turtle, but how do we take 20 years of information from multiple animals and construct an overall behavioral concept for the species? I really enjoy that aspect of the research as well. It’s like a forensic game or a mystery; you get to recreate the past life of these animals and then try to figure out what they were doing and eating (and why) and when they might move from one habitat to another. I love it!
Jeff Seminoff mentioned research techniques such as DNA methylation and telomere research as a potential next step to learn more about these sea turtles. Do you have any plans to investigate that in the near future?
Telomere technology has been explored quite a bit, but the work on sea turtles hasn’t panned out so far. Telomeres are the protective caps on the ends of chromosomes. Telomeres shorten over time, so they’ve been suggested as a way to determine sea turtle age. Methylation seems to have a lot of promise; it has proven to be useful in other taxa like mammals—mostly humans so far—but hasn’t been tried for sea turtles yet. As an animal gets older, more methyl groups attach to its DNA, so we could look at how that changes over time and use it as an indicator for age.
I think it would be fascinating to explore, especially to be able to use skeletochronology in tandem, to help validate this approach to estimating age. We recently had an event with the Southwest Fisheries Science Center that allowed researchers from different divisions to come together. A lot of the tools we have available can cross over to other species and many species have tissues that essentially record time because layers form over time. It’s not just sea turtles – studying layers in teeth from mammals is very common. I’ve done some work on northern fur seal teeth, for example, and fish otoliths (ear bones) are another common example. After the event, people came up to me to discuss methylation and potential avenues to explore this research across species groups. Stay tuned!
Are you working with any partners to advance this research? What are some other aspects of this research that you’d like to investigate in greater detail?

Hawksbill sea turtle.
We have a few collaborations in the works now. Eastern Pacific hawksbill turtles are one of the species we’re studying next, and collaborators in El Salvador have been collecting bones and tissue samples for this project. We’ve also got a large collection of bones from green sea turtle that stranded in along the U.S. West Coast, and we are using some of these approaches to study the life history of both of these turtle populations. We’re also collaborating with folks in Australia to study flatback sea turtles (Larisa is involved in that study, too). Flatbacks are not a U.S. species but this is the first time anyone has done age estimations using skeletochronology on flatbacks. Testing our techniques on other species helps us advance our work for U.S. sea turtles, too.
Also, all of the isotope work we’ve done on turtle bones so far is bulk isotope analysis (looking at all of the carbon or nitrogen in a bone layer). These isotopes can be very indicative of the turtle’s environment and diet. For example, an area with lots of upwelling or nutrients would likely correspond with a wide bone growth layer when the turtle ate well and therefore grew more. Different environmental oscillations can impact the productivity of ecosystems in these Pacific areas also, so that’s something I’d like to investigate soon. Here, we’d be looking at varying isotope values of nitrogen and carbon during El Niño years (where we would expect to see declines in growth) and comparing those patterns with years that have more upwelling. Also, I’d like to investigate how the 2014–2016 Pacific “blob” of warm water affected sea turtles and see if its effects showed up in their bones. We have a good sample set from our U.S. green turtles to test that hypothesis.
The other aspect of this research going forward is using compound specific isotope analysis, whereby we can look at the carbon and nitrogen of individual amino acids in the bone’s different growth layers to get a proxy for the base of the food chain in a particular area, as well as the animal’s trophic position. This would allow us to track changes in the system over 20 years, likely even longer, and then we can incorporate that information into other oceanographic models.
What is the biggest threat or concern for U.S. sea turtle populations right now? What can the public do to help protect sea turtles?
This is why I love the work I do with sea turtles. It comes full circle back to education and outreach.
Fisheries can have a significant impact on these turtles. Be as smart as you can about the seafood you choose. We are lucky, here in the United States, that there has been a lot of work to modify fishing gear, nets, and when and where to fish, and this has really helped sea turtle populations. But the oceans keep changing, so it’s an ongoing process that’s constantly being refined and improved.
Second, pollution in the streets goes right into the ocean. You wouldn’t believe some of the stuff we find in the guts of stranded sea turtles. Plastic bags look like jellyfish, which these turtles love to eat. It is important to keep your trash off the ground, reduce, and recycle.
Finally, climate change is already having a significant effect on sea turtles—temperature changes are skewing the number of males vs. females being born at particular beaches. Nesting beach habitat has also been lost due to rising sea levels and coastal development.